 Preferential usage of particular α- and β-chain genes was similar between all HDs ( Supplementary Fig. TCR-LA-MC PCR sequencing allowed a detailed, unbiased representation of functional TCR gene families, pseudogenes and open reading frame regions 16 ( Supplementary Table 4). 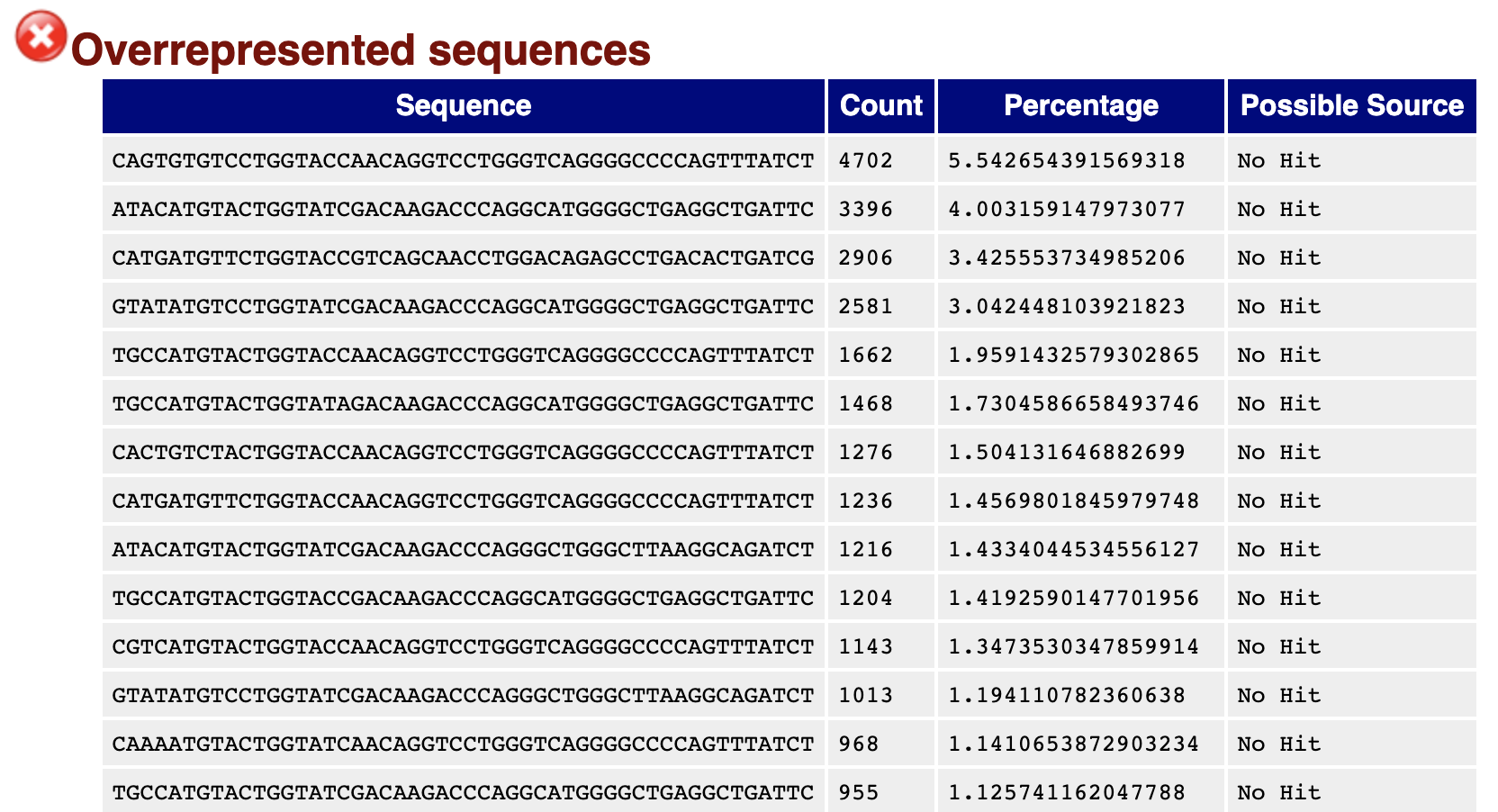
TCR-LA-MC PCR results were consistent with even highly variable sequence numbers, ranging from hundreds to hundreds of thousands ( Supplementary Fig. The 454 sequencing platform was used for HDs 1, 2 and 3 and the MiSeq platform for HDs 4, 5 and 6 ( Supplementary Table 3). To visualize the TCR diversity in HDs with TCR-LA-MC PCR, we first analysed the TCR repertoire in six HDs. Dissection of the TCR repertoire in healthy individuals Furthermore, TCR-LA-MC PCR allows T-cell identification with a resolution capacity of at least 1: 10,000 and down to the single-cell level ( Supplementary Table 2a,b) and readily identifies even rare cells such as invariant natural killer T (iNKT) cells and mucosal associated invariant T-cells 15 ( Supplementary Table 2c). The results showed that even 10 ng of cDNA still provides a reliable representation of the existing TCR repertoire diversity ( Supplementary Fig. We further assessed the sensitivity and applicability of our approach. Using peripheral blood mononuclear cells (PBMC) and sorted T-cell fractions, we first compared available TCR-sequencing methods ( Supplementary Table 1a–e). Here we show the identification of the events leading to the generation of the TCR diversity in healthy donors (HDs), discrimination of different stages of cutaneous T-cell lymphoma in vivo and monitoring of the reactivation of specific TCR clones after in vitro stimulation with peptide pools of two immunodominant cytomegalovirus (CMV) antigens, providing high-resolution characterization of TCR clonality repertoires. To obtain the highest possible resolution at high specificity, we have made use of previously established PCR-based methods: ligation-anchored (LA) PCR 10, 11 and non-restrictive linear amplification-mediated PCR 12, 13, 14, and developed TCR-LA-MC PCR as a new sequencing technology for the immunogenetic characterization of normal and reduced clonality in the α- and β-chain TCR repertoire. Outside of rapid amplification of cDNA ends (RACE) PCR 1, 2, 3, 4, 5, current DNA-based TCR-sequencing technologies require extensive primer multiplexing 6, 7, 8, 9 and are almost exclusively focused on the β-chain repertoire. Comprehensive and unbiased TCR deep sequencing could provide a representative and quantitative repertoire analysis of specific cellular immunity. 
The frequency of a specific CDR3 sequence indicates the abundance of its T-cell clone. The CDR3 region is unique for every T-cell clone and encodes the receptor portion that makes the majority of TCR contacts with antigenic peptides bound by the major histocompatibility complex. Further random insertion and deletion of nucleotides at the rearrangement positions create junctional diversity of the highly variable complementarity-determining region 3 (CDR3 region). These are adjacent to the constant ( C) gene in the heterodimeric α–β or γ–δ TCR. The naive TCR repertoire is assembled in the thymus and formed by somatic rearrangements of non-contiguous genes belonging to the variable ( V), diversity ( D only for the β- and δ-chain) and joining ( J) families (combinatorial diversity). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
